Parameter Selection for HDBSCAN*¶
While the HDBSCAN class has a large number of parameters that can be set on initialization, in practice there are a very small number of parameters that have significant practical effect on clustering. We will first consider those major parameters, and consider how one may go about choosing them effectively. With that out of the way we’ll look at the remaining parameters and see what their effects are – many just effect performance for various different use cases.
Selecting min_cluster_size¶
The primary parameter to effect the resulting clustering is
min_cluster_size. Ideally this is a relatively intuitive parameter
to select – set it to the smallest size grouping that you sih to
consider a cluster. It can have slightly non-obvious effects however.
Let’s consider the digits dataset from sklearn. We can project the data
into two dimensions to visualize it via t-SNE.
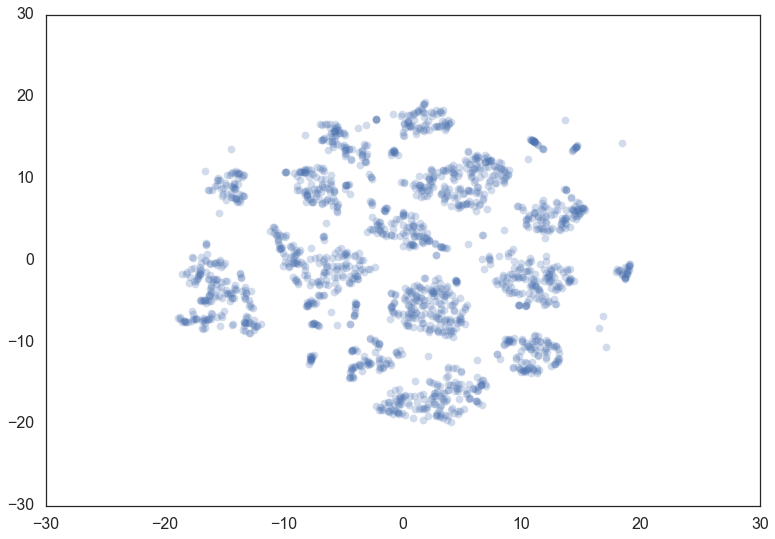
If we cluster this data in the full 64 dimensional space with hdbscan we
can see some effects from varying the min_cluster_size.
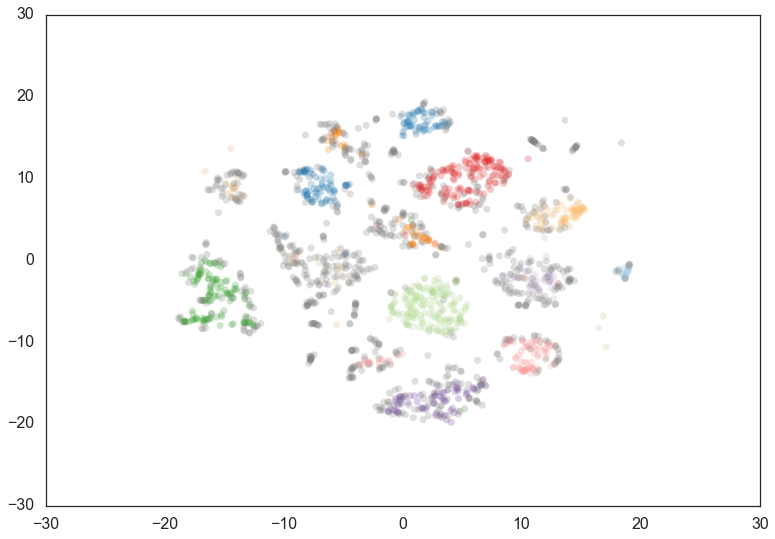
We start with a min_cluster_size of 15.
clusterer = hdbscan.HDBSCAN(min_cluster_size=15).fit(data)
color_palette = sns.color_palette('Paired', 12)
cluster_colors = [color_palette[x] if x >= 0
else (0.5, 0.5, 0.5)
for x in clusterer.labels_]
cluster_member_colors = [sns.desaturate(x, p) for x, p in
zip(cluster_colors, clusterer.probabilities_)]
plt.scatter(*projection.T, s=50, linewidth=0, c=cluster_member_colors, alpha=0.25)
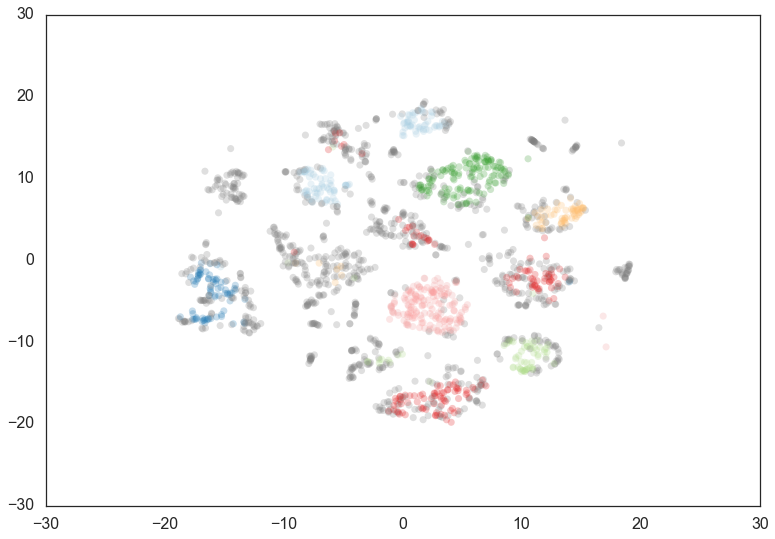
Increasing the min_cluster_size to 30 reduces the number of
clusters, merging some together. This is a result of HDBSCAN*
reoptimizing which flat clustering provides greater stability under a
slightly different notion of what constitutes cluster.
clusterer = hdbscan.HDBSCAN(min_cluster_size=30).fit(data)
color_palette = sns.color_palette('Paired', 12)
cluster_colors = [color_palette[x] if x >= 0
else (0.5, 0.5, 0.5)
for x in clusterer.labels_]
cluster_member_colors = [sns.desaturate(x, p) for x, p in
zip(cluster_colors, clusterer.probabilities_)]
plt.scatter(*projection.T, s=50, linewidth=0, c=cluster_member_colors, alpha=0.25)
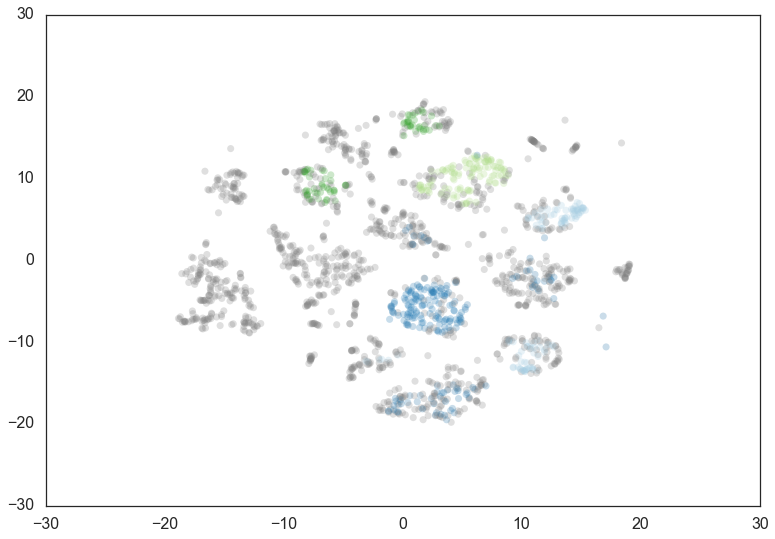
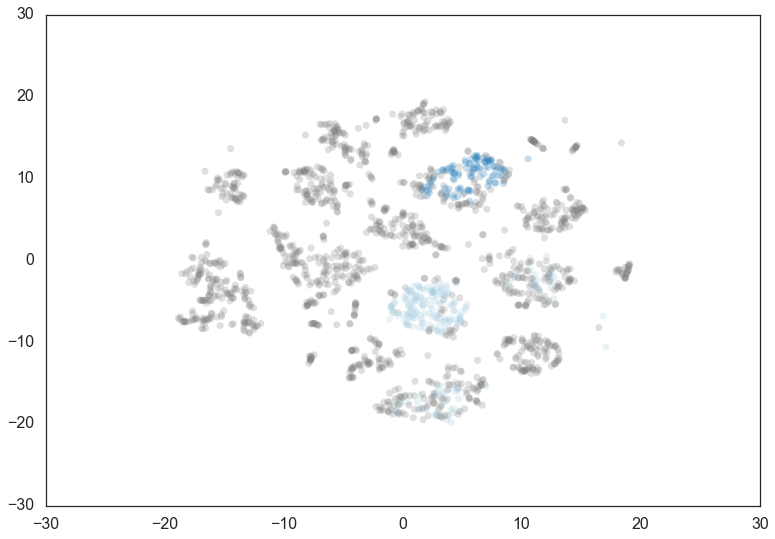
Doubling the min_cluster_size again to 60 gives us just two clusters
– the really core clusters. This is somewhat as expected, but surely
some of the other clusters that we had previously had more than 60
members? Why are they being considered noise? The answer is that
HDBSCAN* has a second parameter min_samples. The implementation
defaults this value (if it is unspecified) to whatever
min_cluster_size is set to. We can recover some of our original
clusters by explicitly providing min_samples at the original value
of 15.
clusterer = hdbscan.HDBSCAN(min_cluster_size=60).fit(data)
color_palette = sns.color_palette('Paired', 12)
cluster_colors = [color_palette[x] if x >= 0
else (0.5, 0.5, 0.5)
for x in clusterer.labels_]
cluster_member_colors = [sns.desaturate(x, p) for x, p in
zip(cluster_colors, clusterer.probabilities_)]
plt.scatter(*projection.T, s=50, linewidth=0, c=cluster_member_colors, alpha=0.25)
clusterer = hdbscan.HDBSCAN(min_cluster_size=60, min_samples=15).fit(data)
color_palette = sns.color_palette('Paired', 12)
cluster_colors = [color_palette[x] if x >= 0
else (0.5, 0.5, 0.5)
for x in clusterer.labels_]
cluster_member_colors = [sns.desaturate(x, p) for x, p in
zip(cluster_colors, clusterer.probabilities_)]
plt.scatter(*projection.T, s=50, linewidth=0, c=cluster_member_colors, alpha=0.25)
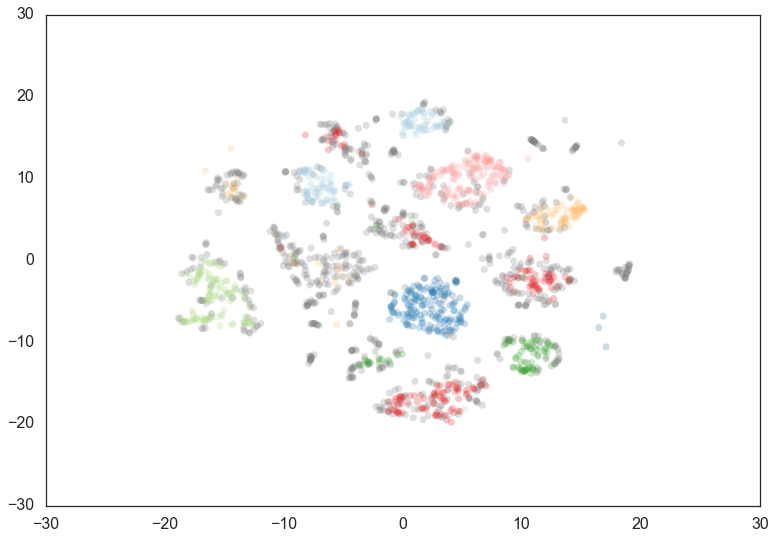
As you can see this results in us recovering something much closer to
our original clustering, only now with some of the smaller clusters
pruned out. Thus min_cluster_size does behave more closely to our
intuitions, but only if we fix min_samples. If you wish to explore
different min_cluster_size settings with a fixed min_samples
value, especially for larger dataset sizes, you can cache the hard
computation, and recompute onlythe relatively cheap flat cluster
extraction using the memory parameter, which makes use of joblib
[link].
Selecting min_samples¶
Since we have seen that min_samples clearly has a dramatic effect on
clustering, the question becomes: how do we select this parameter? The
simplest intuition for what min_samples does is provide a measure of
how conservative you want you clustering to be. The larger the value of
min_samples you provide, the more conservative the clustering –
more points will be declared as noise, and clusters will be restricted
to progressively more dense areas. We can see this in practice by
leaving the min_cluster_size at 60, but reducing min_samples to
1.
clusterer = hdbscan.HDBSCAN(min_cluster_size=60, min_samples=1).fit(data)
color_palette = sns.color_palette('Paired', 12)
cluster_colors = [color_palette[x] if x >= 0
else (0.5, 0.5, 0.5)
for x in clusterer.labels_]
cluster_member_colors = [sns.desaturate(x, p) for x, p in
zip(cluster_colors, clusterer.probabilities_)]
plt.scatter(*projection.T, s=50, linewidth=0, c=cluster_member_colors, alpha=0.25)
<matplotlib.collections.PathCollection at 0x120978438>
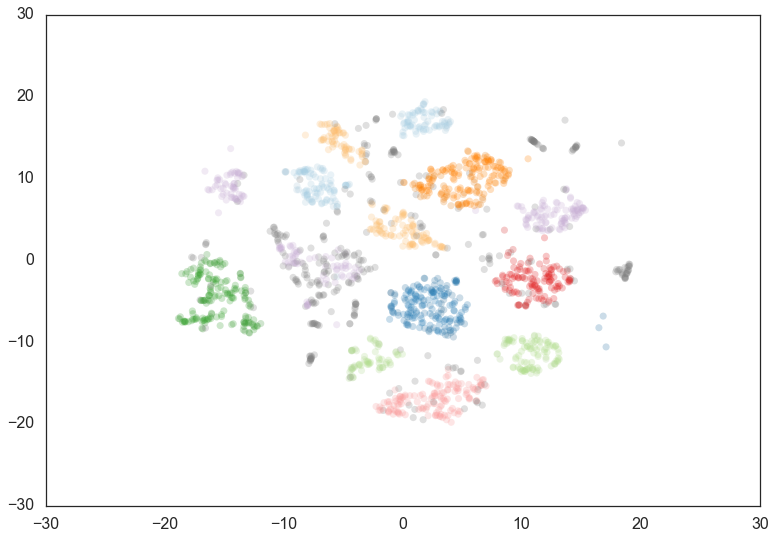
Now most points are clustered, and there are much fewer noise points.
Steadily increasing min_samples will, as we saw in the examples
above, make the clustering progressivly more conservative, culiminating
in the example above where min_samples was set to 60 and we had only
two clusters with most points declared as noise.
Selecting alpha¶
A further parameter that effects the resulting clustering is alpha.
In practice it is best not to mess with this paramter – ultimately it
is part of the RobustSingleLinkage code, but flows naturally into
HDBSCAN*. If, for some reason, min_samples is not providing you
what you need, stop, rethink things, and try again with min_samples.
If you still need to play with another parameter (and you shouldn’t),
then you can try setting alpha. The alpha parameter provides a
slightly different approach to determining how conservative the
clustering is. By default alpha is set to 1.0. Increasing alpha
will make the clustering more conservative, but on a much tighter scale,
as we can see by setting alpha to 1.3.
clusterer = hdbscan.HDBSCAN(min_cluster_size=60, min_samples=15, alpha=1.3).fit(data)
color_palette = sns.color_palette('Paired', 12)
cluster_colors = [color_palette[x] if x >= 0
else (0.5, 0.5, 0.5)
for x in clusterer.labels_]
cluster_member_colors = [sns.desaturate(x, p) for x, p in
zip(cluster_colors, clusterer.probabilities_)]
plt.scatter(*projection.T, s=50, linewidth=0, c=cluster_member_colors, alpha=0.25)